Image credit: Grok (from the provided picture)

HQIV CASP server — Lean ribosome tunnel (default) + optional legacy HKE
Repository: https://github.com/disregardfiat/protein_folder
pyhqiv: https://pypi.org/project/pyhqiv/
Paper (DOI): https://zenodo.org/records/18794890
Lean proofs (hqiv-lean): https://github.com/disregardfiat/hqiv-lean
Paper PDF (GitHub): see hqiv-lean repo (DOI pending on hal.science).
POST / or /predict with sequence(s); GET /help for API details.
Prediction algorithm (default: HQIV Lean ribosome tunnel + bulk solvent)
1) Initial fold (default)
Single chain: co-translational ribosome tunnel with Lean-aligned ε_r(T), pH
screening, dihedral bias toward the HQIV α basin, post-extrusion EM + tree-torque,
then (by default) an HQIV-native sparse OSHoracle quantum pi-phase gate refine on Cα
(Lean ``OSHoracleHQIVNative``; in-code: uses the HQIV-native phase pivot); optional
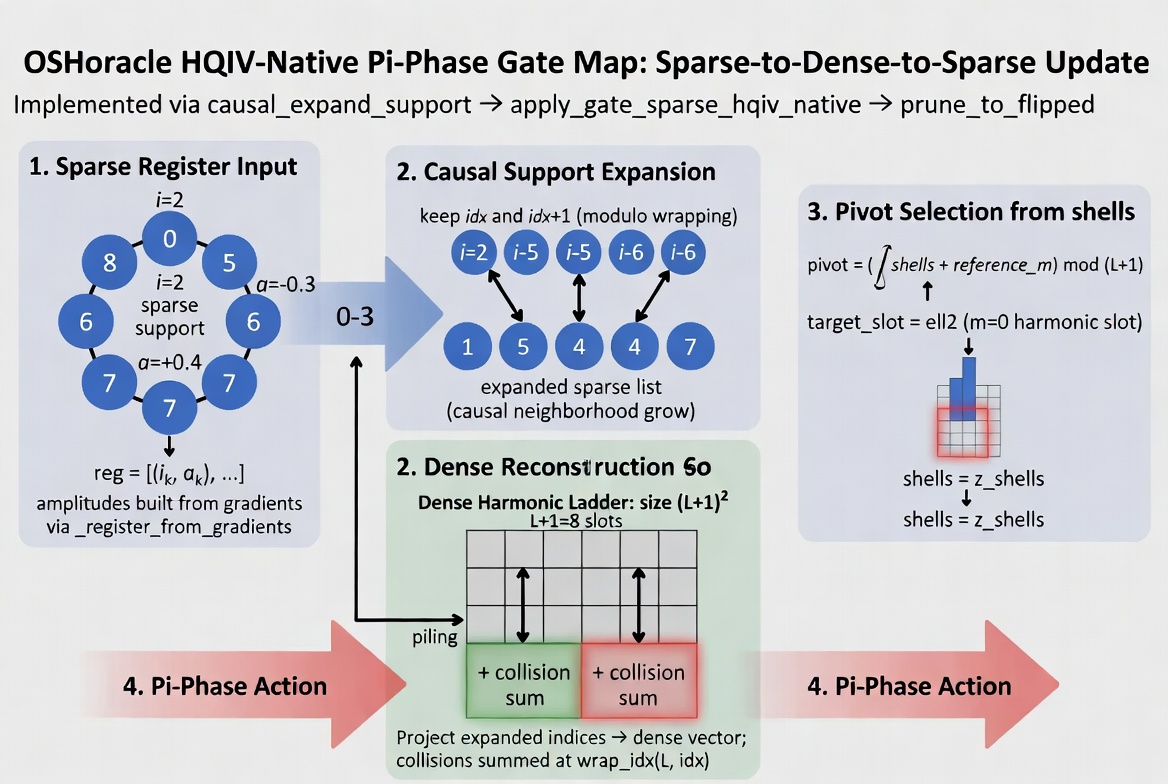
Quantum gate map (HQIV/Lean OSHoracle): causal expand (i->i and i+1), dense harmonic
reconstruction on the harmonic ladder, HQIV-native pi-phase pivot, detect flipped support,
then prune to flipped indices before CA update (only those active residues drive the
coordinate step).
ligands as 6-DOF rigid bodies (included in the tunnel fold for single chain).
Multi-chain: each chain folded the same way, merged with a chain–chain offset; ligands
(if provided) are refined against the **entire** merged complex backbone with the same
Lean bulk-water screening as the fold, then written as HETATM. No inter-chain docking
unless CASP_LEGACY_ASSEMBLY_DOCK=1.
Legacy mode (CASP_LEGACY_HKE_PIPELINE=1): hierarchical HKE + funnel; ligands use
minimize_full_chain; assembly docking as before.
2) Optional tree-torque refinement (CASP_LEAN_TREE_TORQUE_AFTER=1 only)
When set, discrete φ/ψ refinement may run after the Lean backbone (no end flag — run until no allowed moves)
All submissions and retries are piped through discrete φ/ψ refinement with
EM-field-driven unfreezing until convergence (0 accepts in a phase and EM
unfreeze cannot unlock any DOF).
• Discrete DOFs: per-residue φ/ψ in a small set of states (pyhqiv profiles).
• Deterministic downhill: pick the move with lowest ΔE; accept only if ΔE < 0.
• Locking: DOFs that cannot improve are locked; when all are locked, use EM
field torque to decide which to unfreeze and nudge toward a lower-energy
neighbor.
• Tree from the ends: build an EM field from the current backbone; compute
forces; torque τ_i = (r_i - r_anchor) × F_i. When the N- and C-termini are
"free" (not wrapped into the center), anchors are at both ends; torques are
combined per residue; residues to unfreeze are ordered from the ends inward
(min(i, n_res-1-i)). When either terminus is buried (within 0.5 R_g of
center of mass), switch to a single center-of-mass anchor so the torque
tree still gives appropriate translations.
• Re-draw the field: after each unfreeze/nudge, rebuild the backbone from
the current φ/ψ state and recompute the EM field; iterate until no residue
is above the torque threshold (or cap per phase).
• Assembly (A+B, (A+B)+C | ligand): same algorithm, but we add vectors from
residues further from center of mass first (COM anchor, order by distance
from COM descending). This keeps the EM field and connection-point logic
while stepping through refinement from the surface inward.
3) Output
Final PDB is the converged backbone (and ligands if provided). No end flag:
refinement runs until no allowed translations remain.